Just over a year ago, I embarked on a PhD with the LIDo programme here in London. In one of the rarest cases of Twitter being useful, a former colleague posted the call for candidates and I figured I'd apply. Just a few weeks ago, I passed my upgrade here at King's College London, and now I'm heading on the real core of the degree. I wanted to share some thoughts I've had, in the hope they may be useful for others.

Most days I get up, do the morning routine, which occasionally includes setting another neural network running, then cycle into the office. Cycling in London is not for the faint hearted and having a good set of waterproofs is a must! Keeping it chill and not going crazy is the first step to not getting smushed on the roads. Cycle superhighway 2 is my main route, going through Whitechapel, with a left turn past the monument to the Great Fire of London, then over the bridge to Guys. A good ride most days, though you do occasionally get aggro from car drivers.
Checking results of the previous experiments and figuring out what the next ones should be takes a bit of time. This is perhaps where the real science lies, at least at the moment. A large part of this process involves writing code, often beginning with python, then moving over to Rust for speed. Thinking about what has gone right and what has gone wrong, and the path to take is the really tricky part. It's partly intuition, part logic and large parts grumbling and not-quite-getting it.
I'll go to the gym in the morning. Being desk based for most of day, I think it's important to make sure your body is kept fit. The idea that the mind and the brain are separate things was made up by the ancient Greeks (I believe? Happy to be corrected) and has largely been debunked by science. After that, it's back to the office for a shower and then I'm at my desk.
Quite often, there will be emails. Lots of things appear such as ACM news, department events and what not. Supervisors often send things to me that require a bit more attention. It's typically not so bad, despite having two accounts from two different universities to look at. Sometimes there are forms that need filling in. King's makes a big deal about checking up on students every few months to make sure they are doing alright.
There are a lot of events and seminars that take place. Most of the subjects I don't understand at all, but occasionally, I have to talk about my work, either with the department, my research group, or the LIDo programme at large. There's been quite a few of these recently. Though I'm not a huge fan of talking about my work in public, it's nice to get advice and feedback. I think this is a part of science that never gets talked about. We often think of the lonely scientist, striving away at all hours, but it's more common (I'd say) that scientists collaborate and bounce ideas off each other, especially when there is coffee about!
Reading is a big part of the game; I don't do anywhere near enough. Checking what has been published on PLOS One and PLOS Computational Biology gives me some overview. Other places, like OpenAI, Google AI blog occasionally have related articles. PubMed is a source I'm trying to get into. Google Scholar is quite the venerable source of information too, though often people will send me things they think I'll like. Stepping away from the keyboard and actually doing some reading is more difficult than I thought.
When the day is done, I'll ride on back home. I'm a big believer in not reading work emails, or doing serious code commits once I've left the office. It can be hard sometimes, and there are occasions where I'll need to just set a net going at some hour in the evening. Nevertheless, it's good to keep the separation. It's not uncommon though, to get ideas and thoughts about the research when I'm at home. When new ideas hit, I'll write them down and try them later. It's good practice, I think, to give your brain a bit of space. That way, the creative juices can flow.
My background is mostly in programming and computer science, so coming into A.I. and biology means I'm diving into two deep-ends at the same time. Though I did do a bit of work on A.I the year before, I'm certainly no expert. I'm fortunate that I know enough to get started, with the help of my supervisors who've done some work in this area.
Biology is an incredibly large subject. I'd wager bigger than physics and chemistry combined! That's perhaps not entirely fair, because both physics and chemistry feature heavily in biology. My lack of chemistry knowledge was a bit of a stumbling block when I first started on this path. Increasingly though, computer science has really become a major player in this arena. A good example is Deep Mind's Alpha Fold. But computers have been heavily involved in biology for a long time, especially in areas such as X-ray crystallography to name just one.
Most of my work is computational, but my office is next door to some serious biology labs. I have some ideas of what is going on, but I couldn't tell you what half of the machines and tools do, not being a biologist. You get the picture - freezers, pipettes, flasks, microscopes, stirrers, autoclaves, that sort of thing. At some point I'll be making the move over to that side of the corridor, where things get a bit less silicon based, and a bit more carbon based.
I get the impression a lot of computer scientists are coming over to biology. When I was a research software engineer, a number of nasty programs from biologists would appear on my desk. I wondered what was going on because there is really good stuff out there. I jumped the fence to go and have a look and I believe the reason is that biology is incredibly complex, very messy and generates huge amounts of heterogenous data. Cleaning up the data is probably about 90% of the battle. Such efforts tend to result in long pipelines of linked programs, often loosely stitched together with sellotape and string. It makes it hard to optimise and debug. Either that, or everything is done in MATLAB, which is slow and expensive.
Computer vision is one area of CS that has been applied to biology with great success, and since A.I. has revolutionised computer vision, so it will have a big effect on biology. Healthcare in particular is benefitting from the new technology. Examples include Predicting human protein function and Recognising promoters, to name just two I found in a 10 second search. The area has exploded with papers and articles in the last few years. It's an exciting place to be.
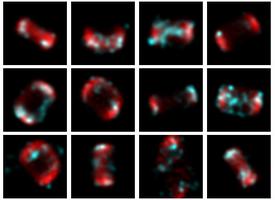
This year, revolves around recreating 3D structure and orientation from 2D images taken with a microscope, with the aid of A.I. So far, it seems to be working, though we've yet to try with the real scary data - that experiment is taking place as I write this! It's taken a lot of experiments to get to this point. This is only the first sub-problem, if you will. These tools will allow us to move on to the next, larger problem.
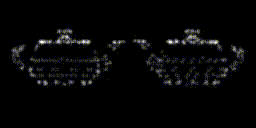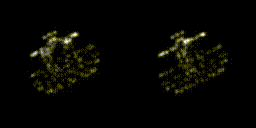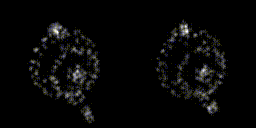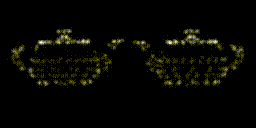
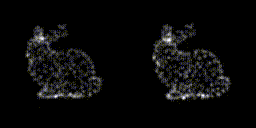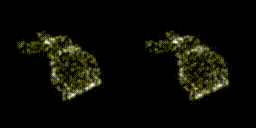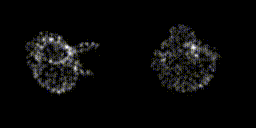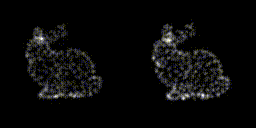
The first year has a lot of getting your head around the general problem space, reading up on the background, getting hold of data, throwing that away, getting more. There's a sense of figuring out which paths might be the most promising, drawing a border around the general problem area before really delving into things in the later years. There are talks to give, lectures to go to, and courses to take as well. I had an online maths course and a basic biology course to complete in addition to starting the research. The LIDo are very keen on employability too, so there is training around associated skills such as presenting work, communicating science and even entrepreneurship.
A large part of this year has been gathering evidence to support our conclusion that this network is actually working as expected. Rigour is the key term here. I spent a good number of weeks trying to get any sort of meaningful statistics out of the data. I tried various approaches before I found one that worked. My supervisor was quite supportive; we need to present an argument and back it up with solid data. One of the problems was our sampling rate. We simply weren't sampling often enough, so we didn't see any changes. It's one of these simple things, but it can make all the difference. It pays to check.
In short, freedom, doing good, working on problems that matter and being treated well. I genuinely believe in science as a noble endeavour (we can talk about how accurate that is in another post) and something worthwhile doing that is bigger than I am. It's also quite freeing in a world where we are assessed, watched, and evaluated 24/7. You are treated like an adult and while that can sometimes go wrong (I have first hand experience of this), it's quite refreshing.
Small things really add up to make for a very lovely life experience. For instance, my office is near Borough Market, so the local food is pretty good, but even the university canteen is great! Showers are provided so I can cycle into work, and there are plenty of clubs, societies and cheap gym memberships. Ultimately, university life at this level feels pretty real and adult, as oppose to startup culture, whose offices feel like creches for man-children, where you are worked to death.
You aren't paid very much, and on that topic, I consider this less of a studentship and more a of job. In fact, one of the top tips I'd give anyone doing a PhD is to treat it as such. Speaking of tips...
I hope the following points might be of use to other students and folks in similar situations. I'm attempting to distill my experience into something useful but it might not be correct. Hopefully it will be helpful to some.
I decided to not only find a topic that was interesting, that I felt I could tackle, but also find a supervisor I could work with. I can't stress this enough! You do have to put a lot of faith in your supervisors; it's very difficult to succeed if you don't have a good working relationship. I spoke with more than 8 different groups before I decided on the combination of project and people that felt right to me.
Backing up your data is really important, but I think it's also important to keep track of how you created the results and data and be able to reproduce them easily. I wrote a bash script that won't let me run an experiment if there are any outstanding files that need to be committed to git. In addition, the code version and all command line parameters are recorded in a file so I can re-run the code with the same settings at any time. I highly recommend this approach. I've included a sample of this script here. This little snippet of bash won't let me start a net training if it's had uncommitted changes:
#!/bin/bash
RED='\033[0;31m'
NC='\033[0m' # No Colour
echo -e "\U1F3CB " ${RED}TRAINING${NC} "\U1F3CB"
if ! git diff-files --quiet --ignore-submodules -- ; then
echo "You have changes that have not been committed. Please commit before continuing."
exit 1
fi
One thing I decided to do early on was to record a lot of statistics as the network trained, so we could see what was happening more clearly. It's proved to be very helpful, so to anyone working on A.I. I'd say visualise early and often. I went so far as to build a website to house all the results - this way, I could share my results with my supervisors easily. I have a single script that generates a whole slew of graphs and images, then uploads them for viewing.
Folks say you should write and write early. I've kept a diary and various notes though I'm sure there's more I should be doing. I use Zotero to keep track of everything I read and why. Hoping this will pay off when I come to write up.
A large part of a PhD seems to be project management. It's really easy to go down a rabbit hole, getting a thing to work and lose all sense of time and proportion. Sometimes, things just won't work and you need to move on, albeit with a slightly different path. It's somewhat in contest with the rigour point I made earlier. I suppose that's part of the skill of doing science.
Meeting with others, sharing your ideas and talking to others about their research is a good thing to do. It's emphasised a lot in the school I'm in. There are many seminars, whatsapp groups and similar social networks for all sorts of things. Though I don't go to them all (it's important to develop a good filter for these things), going to a few really helps fuel up the mental batteries, prompts new ideas and stops you from feeling isolated.
I'd also say, read a bit of philosophy of science. It had quite an impact on me and my view of how science is done and what it means. I'm still working on this part. It's not easy reading at times but I think it's something that isn't often mentioned these days, but seems quite important to me.
I've had doubts and fears during this first year. I think that might be normal, or at least more common than folks let on. The problems PhD students work on are hard. There's no getting around that. It's much more like a marathon than a sprint; there will be many ups and downs. It can be hard to smooth over these and surf to the end. For some, it might not be the right time. I'm very fortunate that I have this opportunity. I suspect I am, on average, about 15 years older than most of the students in my cohort.
I think it's worth saying doing a PhD won't massively change you; you don't start at one end as a naive student and come out the other as a super smart doctor. It's also not guaranteed that anything you discover will make a huge dent in your discipline, but that's fine. It's a series of events that are worth savouring, enjoying or gritting your teeth and getting through. Though hopefully, by the end, I'll at least be a little wiser and have helped prune humanities web of knowledge, if just a little.
